Multiple Myeloma Network Portal
Welcome to Multiple Myeloma Network Portal to explore causal mechanistic regulatory network and programs for multiple myeloma.
Search the Model
Please use the search box to query genes, regulators, regulons, or mutations.
Search for genes or regulators from the Multiple Myeloma Regulatory Network model by using autocomplete in the search field based on your input. For example input IRF and it brings up all the possible matches. Then select the correct search term from the drop-down list and click the Search button.
Examples:
regulator=E2F1
gene=BRCA1
Explore the Model
We developed the MINER pipeline to infer transcriptional regulatory networks from gene expression data and apply them to the characterization and prediction of phenotypes. MINER builds upon our previous work with the SYstems Genetics Network AnaLysis (SYGNAL) pipeline insofar as it enables the same core functionalities of mechanistic and causal inference, but does so with a new suite of algorithms that enable new applications in the network-based prediction of clinical outcomes.
Application of MINER to the pre-processed MMRF IA12 data successfully generated a Causal Mechanistic (CM) TRN of MM. The network features 15,192 genes partitioned into 1,233 co-expression clusters (i.e., without inferred co-regulation), 8,549 genes partitioned into 3,203 co-regulated modules (called regulons herein) that are regulated by 392 unique transcription factors, and 124 causal drivers, including somatic mutations, translocations, and cytogenetic abnormalities. In total, the MINER network comprises 13,587 unique CM flows – links from mutations to regulators to co-regulated genes.
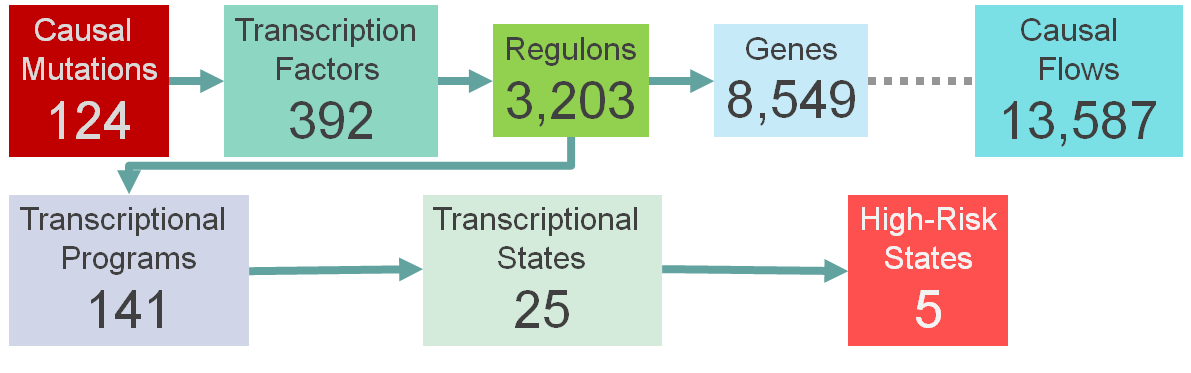
MINER Model Overview

Please use the search box to query genes, regulators, regulons, or mutations.
Explore Causal Mechanistic Flows
13,587 causal mechanistic flows
Explore table of influences from mutations to regulators to regulons
Causal Mechanistic Flows (CM Flows) are statistically significant links between putative causal events (somatic mutations, copy number variations, chromosomal translocations, etc.) and the activity levels of regulators and regulons. In the causal flow, a mutation may causally activate or deactivate a downstream regulator which then might up- or down-regulate a regulon that contains genes with similar expression profiles and binding sites.
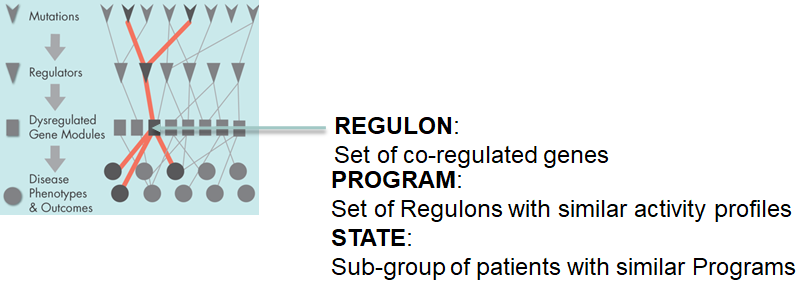

CM Flows

Literature Validations
Explore the literature evidences for causal associations between different components.
We performed comprehensive literature validation for multiple associations included in the causal mechanistic flows. We provide easy to navigate visualization app to explore these validations.
Model Overview
| # | Description |
|---|---|
| Regulons | |
| Mutations | |
| Regulators | |
| CM Flows | |
| Transcriptional Programs | |
| 881 | Patients |

